About our group
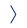
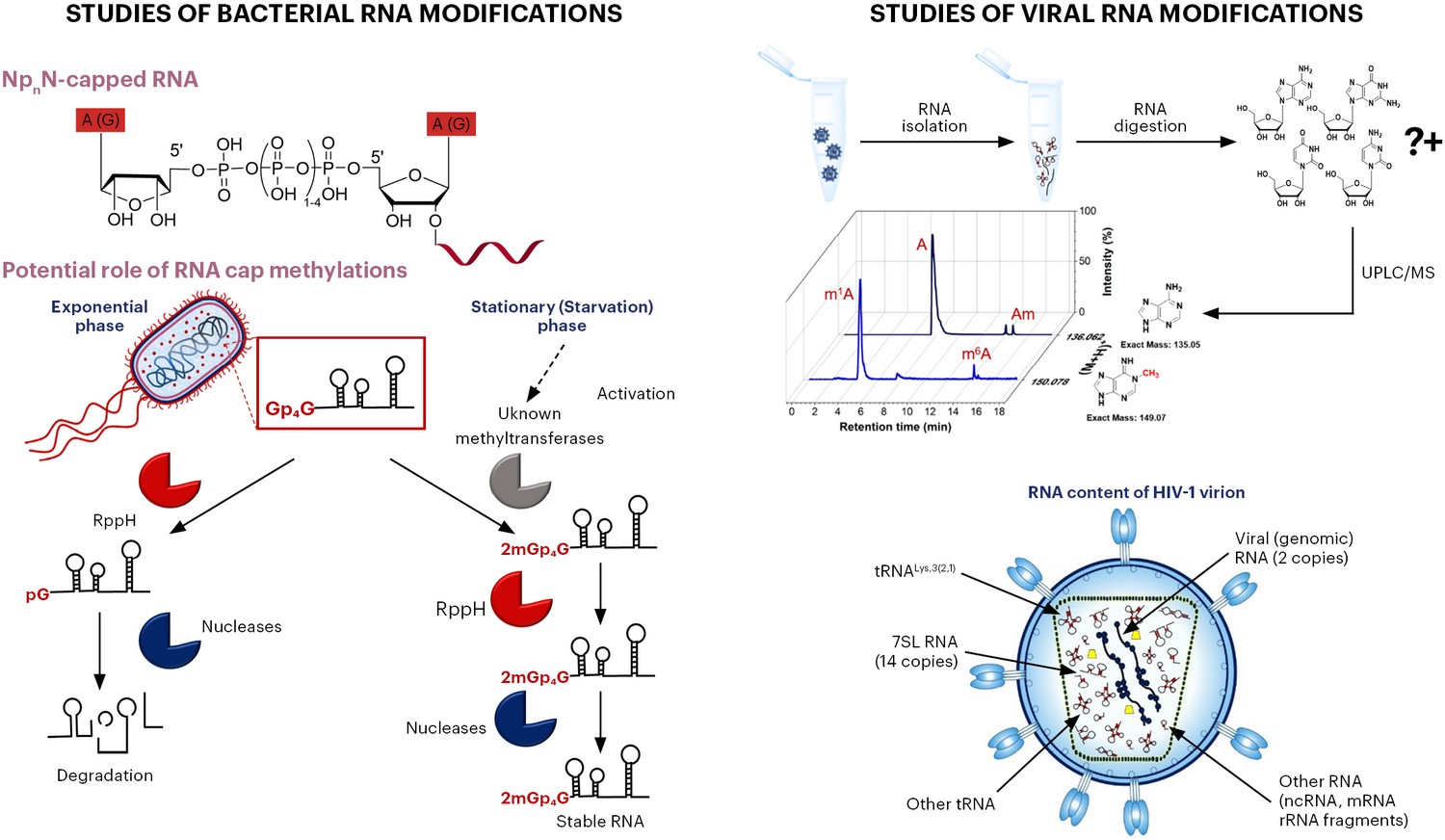
The target of our group is to reveal new function of non-coding mainly regulatory RNAs. We believe that chemical RNA modifications are responsible for the vast majority of various RNA functions. Only few of them have been so far discovered, mainly because of the lack of sensitive methods. Recently, UPLC/MS was used for discovery of two new RNA modifications NAD and CoA in prokaryotes. Thanks to NAD captureSeq it was found that NAD is covalently attached to 5´end of some regulatory RNAs and short fragments of certain mRNAs in prokaryotes and eukaryotes. It was also found that such RNA is more stable and less prone to degradation by exonucleases. We suggest that viruses are perfect model systems for searching for new RNA modifications, as they have simple intrinsic organization and they are amplified in infected cells. We concentrate on the clinically relevant viral strains (e.g. HIV, Picornaviruses and Vaccinia Virus) and the development of new capturing methods for known RNA modifications of viral or bacterial RNAs as well as on the methylation profiling of viral RNA. In the next phase, we will study function, biogenesis and biodegradation of viral RNA modifications.